Validate the binding strength of individual T cell peptide epitope candidates in order to judge their immunogenic potential.
Peptide Epitope Validation
Accurately determine half maximal inhibitory concentrations (IC50), representative of the binding affinity where your unlabeled test peptide competes with a labeled reference peptide for HLA binding from a wide range of alleles.

Introduction
Peptide epitopes are one of the most concise pieces of information required by the T cells to generate an immune response and play a key role in the development of novel immunotherapies for infectious and malignant diseases. Although various peptide epitope discovery methods are available, the results are often not in accordance with physical binding data and suffer from a significant rate of false-positive and false-negative results. HLA Protein Technologies Inc’s peptide epitope validation services offer solutions to these problems using uniquely efficient means of validating immune target molecules that are potentially recognized by T cells. Validation successfully eliminates false-positive or negative results, confirms the peptide candidate’s binding affinity, and facilitates the proper prioritization of these potential T-cell epitopes according to their greatest potential.
How it Works
Provide a synthetic peptide
Select peptides from previous screening results originally derived from target source proteins in cancer and infectious disease applications, from putative epitopes originated from predictive algorithms or select altered or mutated peptides from viral variance studies or tumor-specific mutations.
Select an HLA molecule
Select from a broad representation of HLA alleles to be applied in our peptide binding assays. Our service is very flexible, ensures expedited results, and employs highly-purified single specificity soluble HLA proteins from mammalian cell lines to guarantee outstanding performance in accuracy, sensitivity and reproducibility.
Receive the data
Inhibitory concentrations are determined by incubating sHLA with a labeled reference peptide in the presence of different concentrations of competitor peptide. We deliver a calculated logIC50 value as measure of the effectiveness of the competing test peptide. Affinity categories will prioritize your logIC50 values into high, medium, or low affinity binders.
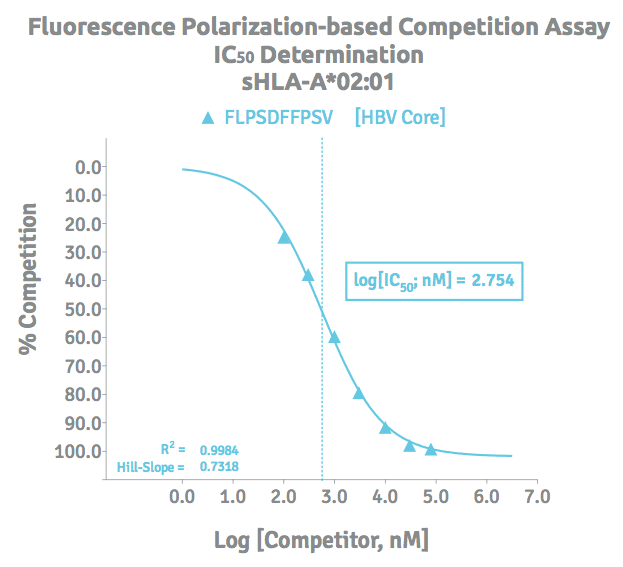
Peptide Binding Assay Results
A logIC50 value is determined by plotting the response values as a function of the logarithms of competitor concentrations and applying a nonlinear regression dose response model with variable slope for analysis.
HLA Validation Service
Download our list of HLA validation services to view all available proteins as well as services in development.
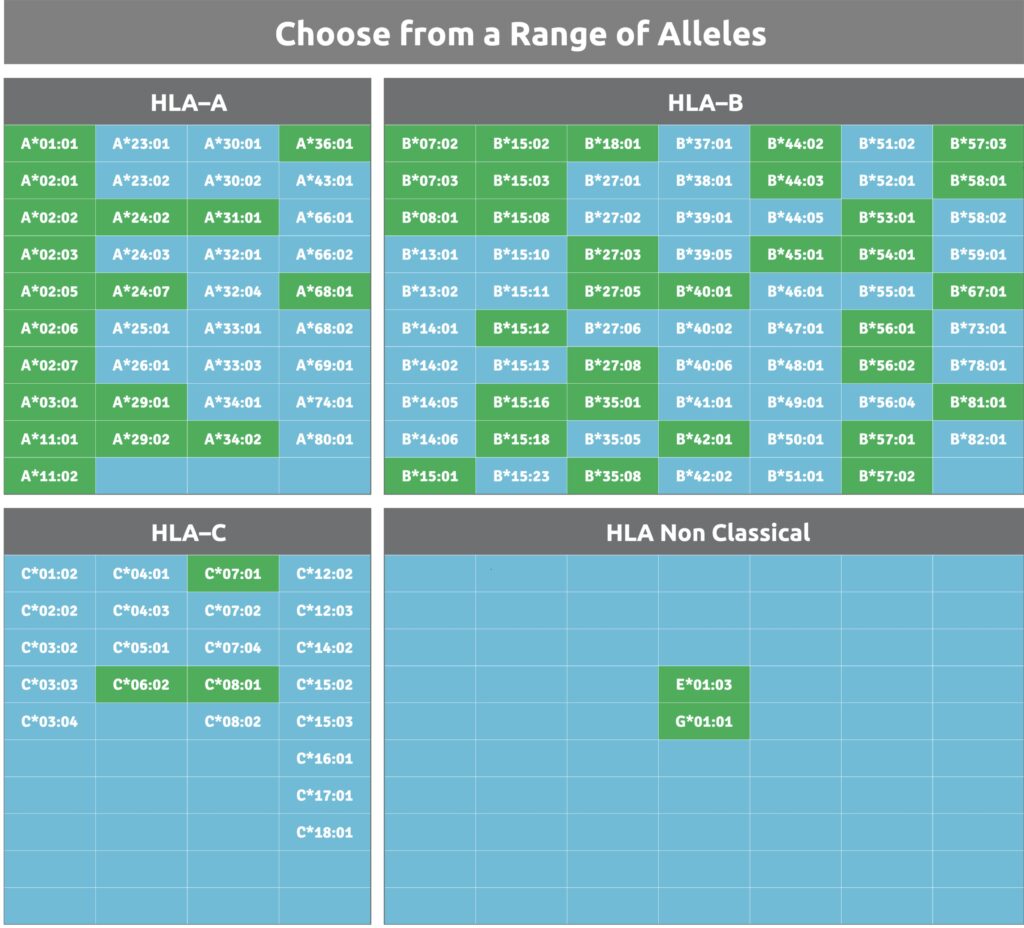
Validation Options
Validate Screening Hits
Confirm the validity of your screening hits and properly prioritize potential T-cell epitopes according to their HLA affinity. Successful elimination of false positive or negative screening results will increase confidence and reduce risks working with your target source proteins from cancer or infectious diseases applications.
Validation of Predicted Peptides
Our HLA binding assays do what prediction tools cannot. They confirm the physical binding characteristics of actual peptides. Directly measure the binding strength of your predicted peptides, allow proper prioritization and successfully eliminate false positive or negative results generated by computer algorithms.
Validate Mutational Variations
Validate neo-epitope candidates from tumor-specific mutations for their increased or decreased binding or peptides covering viral escape mutations measuring the loss of reactivity against the wild-type to determine a potential sequential accumulation of CTL escape in patients during disease progression.
Validate Modifications
Explore whether target modulation will improve your binding potency. Identify vaccine candidates not providing the desired immune response levels and test their redesign by easily alter potential anchor positions for improved binding and quickly assess and compare the results of those modifications.
Competition Binding Assay
HLA binding assays provide uniquely efficient means in validating immune target molecules that are potentially recognized by cytotoxic T cells. We help researchers advance qualified candidates quickly through the development process by facilitating peptide affinity confirmation and ranking based on binding characteristics using a novel state-of-the-art assay procedure, using competition-based fluorescence polarization.
Competition Analytics (LogIC50 –Determination)
The peptide binding assay delivers IC50 determinations, a measure of the effectiveness of a competitor peptide in inhibiting a reference fluorescent-labeled peptide tracer from binding the HLA ligand recognition site. Dose dependent inhibition is a logarithmic phenomenon with a sigmoid function, and as such is the applicable format to view and report the data as logIC50.
LogIC50 Ranking
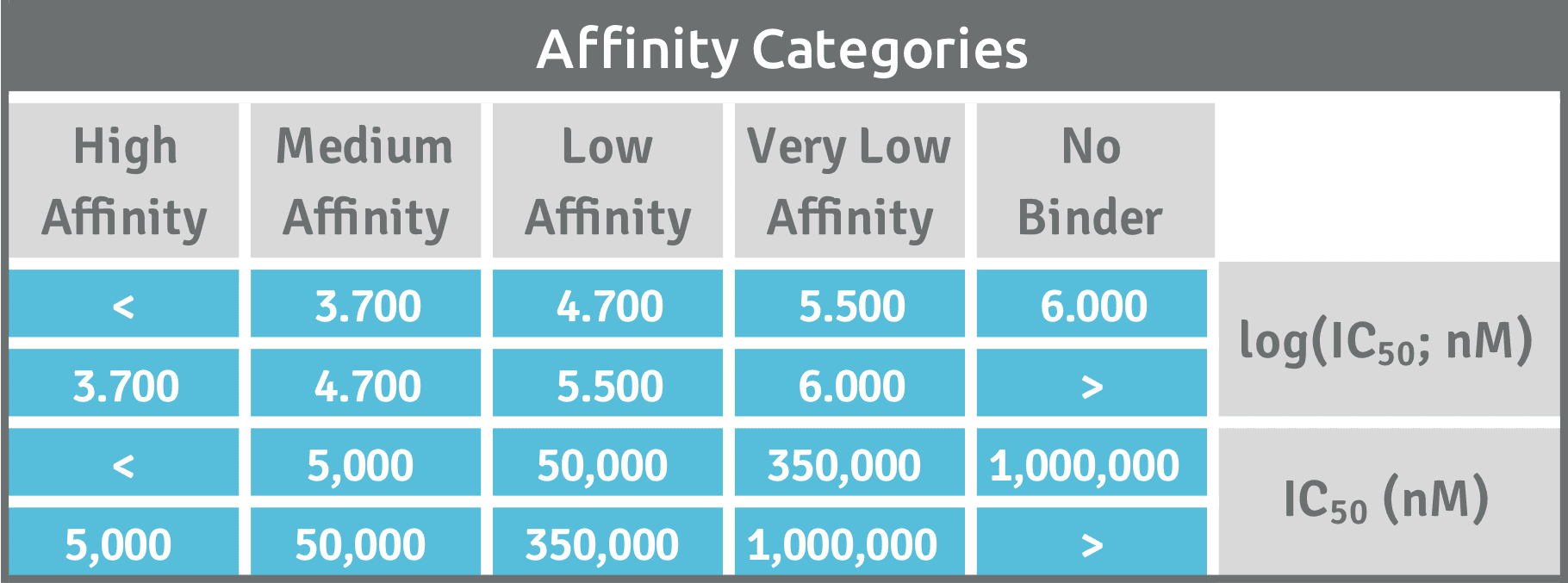
In order to judge the T cell peptide epitope candidate’s immunogenic potential and value in the development of novel immunotherapies, relative affinities of multiple peptide ligands for the same HLA receptor need to be compared. Peptides with higher affinity are more likely to be suitable T cell epitopes and are preferred over the peptides with lower affinity. Our approach ranks the identified peptides to prioritize epitopes with greatest potential based on preset affinity categories. Our classification of the peptide-binding affinity into high (H), medium (M), low (L), and very low (VL) is comparable to the classifications set by other investigators that have used the same reference peptides [Ref].
Frequently Asked Questions
What do I get when I send a peptide for validation?
High affinity binding is the critical factor controlling immunogenicity of peptides. We will determine logIC50 values, provide the associated dose-response curve in graphical display and ranked your candidate into affinity categories of high, medium or low affinity.
What is an IC₅₀?
An IC50 value is the half maximal inhibitory concentration in a dose response model that tells us the concentration at which a competitor peptide is able to inhibit the standard HLA binding process to the reference tracer peptide by 50%.
What is a competition peptide binding assay?
Competition assays are based on peptide binding to sHLA molecules where competition for binding between the test peptide (competitor) of interest and a fluorescein-labeled HLA class I binding peptide (tracer) is used as read out.
Talk to an expert today
Want to know more about our Validation Systems?
If you’re still wondering if our services are the right technology to take you to the next level and are looking for some clarity, reach out to our experts and someone from our team will get back to you in no time.
Expert advice, whenever you need it!
Related Products & Services
Soluble HLA-B
Select from a pool of over 65 HLA-B alleles to drive your immunological research to identify antibody immune responses or visualize antigen-specific immune cells.
Soluble HLA-C
Discover a selection of over 20 HLA-C alleles, the dominant ligand for KIR on NK cells for your immunological research.
T cell Epitope Mapping
Discover your next immunotherapy targets utilizing a high-throughput approach to create individual epitope maps on any chosen protein that is potentially recognized by T cells.

